A force generating living polymer: experiments and theory to understand how it works
 Wednesday, 30th January 2013. 12:00-13:00
Wednesday, 30th January 2013. 12:00-13:00
Marisela Vélez
(Instituto de Catálisis y Petroleoquímica (CSIC) and IMDEA Nanoscience, Madrid)
ABSTRACT:
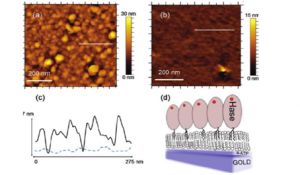
FtsZ is a bacterial cytoskeletal protein that polymerizes on the inner surface of the bacterial membrane and contributes to generate the force needed for cell division. In the presence of soluble modulators the individual protein monomers interact longitudinally to form filaments that can then aggregate to form higher order structures on a surface. This filament aggregates are dynamic and exchange monomers from the solution. The final outcome of this dynamic rearrangement on the surface is the generation of force that bends the cell membrane inward. Experiments on model systems using atomic force microscopy have allowed studying the dynamic behavior of individual filaments and, in combination with theoretical models that can explain the experimental results, we are gaining understanding at the molecular level about the force generation mechanism.
Paez , A., Mateos-Gil, P., Hörger, I., Mingorance, J., Rivas, G., Vicente, M., Vélez, M. and Tarazona, P. “Simple modeling of FtsZ polymers on flat and curved surfaces: correlation with experimental in vitro observations” (2009) PMC Biophysics, 2:8.
Mingorance, J., Rivas, G., Vélez, M., Gómez-Puertas, P., Vicente, M. “Strong FtsZ is with the force: mechanisms to constrict bacteria” Trends in Microbiology, 2010, 18(8) pp. 348 – 356.
Mateos-Gil,P; Marquez, I.; López-Navajas,P.; Jiménez, M.; Rivas,G.; Mingorance, J.; Vicente,M. and Vélez,M. “FtsZ polymers bound to lipid bilayers through ZipA form dynamic two dimensional networks” BBA – Biomembranes (2012) 1818: 806- 813.
Mateos-Gil, P.; Paez, A.; Hörger, I.; Rivas, G.; Vicente,M.; Tarazona, P.; and Vélez, M. “Depolymerization dynamics of individual filaments of bacterial cytoskeletal protein FtsZ” 2012 PNAS 109 (21) 8133-8138.